Mutation in tomato using EMS and gene discovery.pptx
•
0 likes•6 views
Mutationin tomato using EMS and gene discovery
Report
Share
Report
Share
Recommended
Recommended
Agrobacterium-mediated transformation of rough lemon (Citrus jambhiri Lush) w...

Agrobacterium-mediated transformation of rough lemon (Citrus jambhiri Lush) w...Mohammed Naziya kaleem
More Related Content
Similar to Mutation in tomato using EMS and gene discovery.pptx
Agrobacterium-mediated transformation of rough lemon (Citrus jambhiri Lush) w...

Agrobacterium-mediated transformation of rough lemon (Citrus jambhiri Lush) w...Mohammed Naziya kaleem
Similar to Mutation in tomato using EMS and gene discovery.pptx (20)
TILLING & Eco-TILLING : Reverse Genetics Approaches for Crop Improvement

TILLING & Eco-TILLING : Reverse Genetics Approaches for Crop Improvement
Agrobacterium-mediated transformation of rough lemon (Citrus jambhiri Lush) w...

Agrobacterium-mediated transformation of rough lemon (Citrus jambhiri Lush) w...
Mut map genetic mapping and mutant identification without crossing

Mut map genetic mapping and mutant identification without crossing
Green Synthesis of Silver Nanoparticles exhibits Anti-Metastasis and Anti-Bio...

Green Synthesis of Silver Nanoparticles exhibits Anti-Metastasis and Anti-Bio...
Marker assisted selection of male sterility in rice --vipin 

Marker assisted selection of male sterility in rice --vipin
Resistance/tolerance to root-knot nematode Meloidogyne incognita (Kofoid and ...

Resistance/tolerance to root-knot nematode Meloidogyne incognita (Kofoid and ...
Impact of EMS Induction on Morphological, Anatomical and Physiological Traits...

Impact of EMS Induction on Morphological, Anatomical and Physiological Traits...
More from Baban Jeet
More from Baban Jeet (19)
Use of sensor technology for real monitoring quality of vegetable crops

Use of sensor technology for real monitoring quality of vegetable crops
Agronomic biofortification of Iodine and selenium in vegetable crops

Agronomic biofortification of Iodine and selenium in vegetable crops
Capsaicin and Sugar estimation in vegetables .pptx

Capsaicin and Sugar estimation in vegetables .pptx
Introduction to greenhouse equipments, Materials of construction for traditio...

Introduction to greenhouse equipments, Materials of construction for traditio...
Forcing Techniques For Raising Summer Vegetables .pptx

Forcing Techniques For Raising Summer Vegetables .pptx
Seed pelleting scope and constraints in vegetables.pptx

Seed pelleting scope and constraints in vegetables.pptx
Improving nutrient status of vegetables crops through foliar fertilization .pptx

Improving nutrient status of vegetables crops through foliar fertilization .pptx
Development of male sterile lines in brinjal and chilli.pptx

Development of male sterile lines in brinjal and chilli.pptx
Morphologica, anatomical changes during water stress

Morphologica, anatomical changes during water stress
Alkali soils and its management for vegetable crops .pptx

Alkali soils and its management for vegetable crops .pptx
Molecular characterization of human ABHD2 as TAg lipase .pptx

Molecular characterization of human ABHD2 as TAg lipase .pptx
Agroecology for sustainability in vegetable crops .pptx

Agroecology for sustainability in vegetable crops .pptx
present status and prospects of protected cultivation in vegetable crops .pptx

present status and prospects of protected cultivation in vegetable crops .pptx
Recently uploaded
Organic Name Reactions for the students and aspirants of Chemistry12th.pptx

Organic Name Reactions for the students and aspirants of Chemistry12th.pptxVS Mahajan Coaching Centre
“Oh GOSH! Reflecting on Hackteria's Collaborative Practices in a Global Do-It...

“Oh GOSH! Reflecting on Hackteria's Collaborative Practices in a Global Do-It...Marc Dusseiller Dusjagr
Mattingly "AI & Prompt Design: The Basics of Prompt Design"

Mattingly "AI & Prompt Design: The Basics of Prompt Design"National Information Standards Organization (NISO)
Recently uploaded (20)
18-04-UA_REPORT_MEDIALITERAСY_INDEX-DM_23-1-final-eng.pdf

18-04-UA_REPORT_MEDIALITERAСY_INDEX-DM_23-1-final-eng.pdf
Separation of Lanthanides/ Lanthanides and Actinides

Separation of Lanthanides/ Lanthanides and Actinides
Contemporary philippine arts from the regions_PPT_Module_12 [Autosaved] (1).pptx![Contemporary philippine arts from the regions_PPT_Module_12 [Autosaved] (1).pptx](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
![Contemporary philippine arts from the regions_PPT_Module_12 [Autosaved] (1).pptx](data:image/gif;base64,R0lGODlhAQABAIAAAAAAAP///yH5BAEAAAAALAAAAAABAAEAAAIBRAA7)
Contemporary philippine arts from the regions_PPT_Module_12 [Autosaved] (1).pptx
Organic Name Reactions for the students and aspirants of Chemistry12th.pptx

Organic Name Reactions for the students and aspirants of Chemistry12th.pptx
“Oh GOSH! Reflecting on Hackteria's Collaborative Practices in a Global Do-It...

“Oh GOSH! Reflecting on Hackteria's Collaborative Practices in a Global Do-It...
Z Score,T Score, Percential Rank and Box Plot Graph

Z Score,T Score, Percential Rank and Box Plot Graph
Mattingly "AI & Prompt Design: The Basics of Prompt Design"

Mattingly "AI & Prompt Design: The Basics of Prompt Design"
Hybridoma Technology ( Production , Purification , and Application ) 

Hybridoma Technology ( Production , Purification , and Application )
Privatization and Disinvestment - Meaning, Objectives, Advantages and Disadva...

Privatization and Disinvestment - Meaning, Objectives, Advantages and Disadva...
Beyond the EU: DORA and NIS 2 Directive's Global Impact

Beyond the EU: DORA and NIS 2 Directive's Global Impact
Introduction to ArtificiaI Intelligence in Higher Education

Introduction to ArtificiaI Intelligence in Higher Education
Mutation in tomato using EMS and gene discovery.pptx
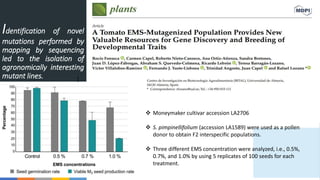
- 1. Identification of novel mutations performed by mapping by sequencing led to the isolation of agronomically interesting mutant lines. ( Moneymaker cultivar accession LA2706 S. pimpinellifolium (accession LA1589) were used as a pollen donor to obtain F2 interspecific populations. Three different EMS concentration were analyzed, i.e., 0.5%, 0.7%, and 1.0% by using 5 replicates of 100 seeds for each treatment.
- 3. Total no of 16,104 MM seeds were EMS treated 12,560 M1 plants were obtained, from which 7379 (58.75%) produced M2 segregating progenies. M2 progenis were screened under greenhouse conditions along eight crop cycles to detect recessive mutants 2800 mutants (37.95%) were detected and grouped into 14 phenotypic categories
- 4. EMS mutant lines displaying alterations during vegetative and reproductive development
- 5. 5 Gene isolation of casual mutation Scanning electron microscopy (SEM) proved that sepals of mutant plants lack the abscission zone between each sepal. Therefore, they have called this mutant sepal indehiscent (sin).
- 7. Tomato mutants have been and are still directly used in breeding programs, but natural variation of agronomically interesting alleles is scarce in natural germplasm. Induced mutagenesis rises as one of the most relevant genomics tools to allow for the functional characterization of most of the unknown tomato genes. The identification of novel mutations performed by mapping by sequencing has led to the isolation of causal mutations of agronomically interesting mutant lines. Such is the case of the line UAL-7334 (sin), where this approach has allowed us to relate a single nucleotide deletion in the Auxin Response Factor 10A (SlARF10A) gene to this mutant phenotype. The use of another species genetically distant from cultivated tomato provides a huge number of polymorphisms for linkage analysis, simplifying the bioinformatic workflow, and ensures a high mapping accuracy. Conclusion
Editor's Notes
- © Copyright Showeet.com – Free PowerPoint Templates
- © Copyright Showeet.com – Free PowerPoint Templates